1 引言
学术论文是科研人员进行研究发现发表和学术观点交流的主要形式[1]。在互联网时代,学术论文的发表和传播普遍采用电子化媒介,科研人员可查阅到海量的学术论文。这样的科研模式提出了高效过滤和梳理海量的学术文献知识的需求。因此,需要解决从非结构化的学术文献文本中进行信息提取和知识组织的问题。
深度学习算法具备从数据中学习深层抽象特征的能力,在分类任务中免去繁琐的特征构建步骤。本研究采用深度学习语言表征模型,改进模型输入,以标注数据集对模型参数进行调优,训练获得学术论文语步结构分类深度学习模型。
2 相关研究
图1
机器学习语步分类研究中的一个关键步骤是分类算法输入特征的构建。从特征构建的角度分析,可分为以下类型。
相比而言,句子粒度的深度学习语言表征模型,如BERT(Bidirectional Encoder Representations from Transformers)[24],可训练获得句子粒度的向量表达,且蕴含了句子的潜在语言学特征,可根据特定任务进行参数调优。基于此,本文受BERT语言表达模型的启发,以学术论文训练的BERT模型参数为基础改进模型输入,增加句子位置向量,训练论文语句的向量表达,输入分类器实现论文语步结构分类。此方法对基于海量文本训练的深度学习语言表征模型进行迁移学习,调节模型参数以学习学术文献的语言特征,既提高了模型的训练速度,又能适应学术论文的应用场景,实现算法运行速度和效果的提升。
3 深度学习语步分类方法
图2
图2
语步分类深度学习模型结构
Fig. 2
Deep Learning Classification Model Structure for Argumentative Zoning
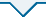
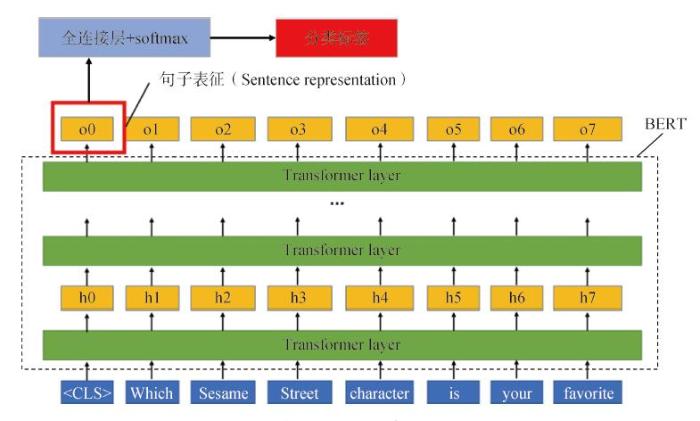
BERT模型通过Transformer堆栈构建双向编码器表征。模型训练方法是采用大型的语料库,对语料进行随机屏蔽,并训练预测屏蔽内容,直至模型的损失函数最小。BERT通过联合调节所有层中的上下文预先训练深度双向表征,对上下文语境有较强的学习能力,具体训练机制是从语料文本中随机抽取15%单词,通过BERT模型预测这些被抽取单词位置的内容。BERT的双向机制在处理一个词的时候,能学习到该词前文和后文单词的信息,从而学习上下文的语境信息。
模型的输入以句子为单位,并在每个句子前加上一个特别的分类嵌入<CLS>,作为输入系列的第一个Token。对于句子分类任务,模型学习到的最后一层隐含状态的第一个位置的输出即作为句子的向量表达。
3.1 改进模型数据输入
学术论文写作遵循相对固定的章节顺序,一般写作顺序为:研究背景,相关研究,方法,结果,讨论,结论。句子在论文中出现的位置可为推断句子语步属性提供有用信息。例如,句子出现在论文开始部分,则大概率属于研究背景。基于此,本文设计改进了BERT模型的输入,增加表征输入句子在篇章中所处位置的向量,与原有的输入嵌入向量进行加和作为模型的输入。
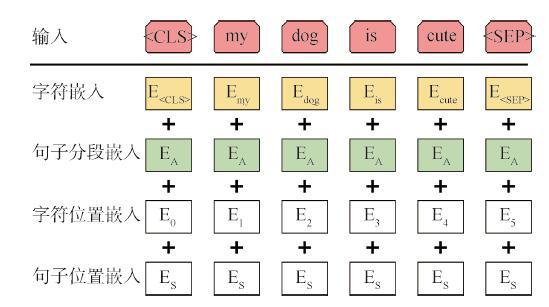
BERT模型的输入是通过三种嵌入向量加和而成,分别为字符嵌入(Token Embeddings)、句子分段嵌入(Segment Embeddings)、字符位置嵌入(Position Embeddings)。模型在训练过程中构建三种嵌入向量的查询表,并将其作为模型参数在训练过程中学习。如图3所示,类似于字符位置嵌入向量,本文在此基础上提出非学习输入向量——句子位置嵌入向量,表征当前输入句子在论文中的位置,即本研究模型的输入为上述4种向量嵌入的加和。
图3
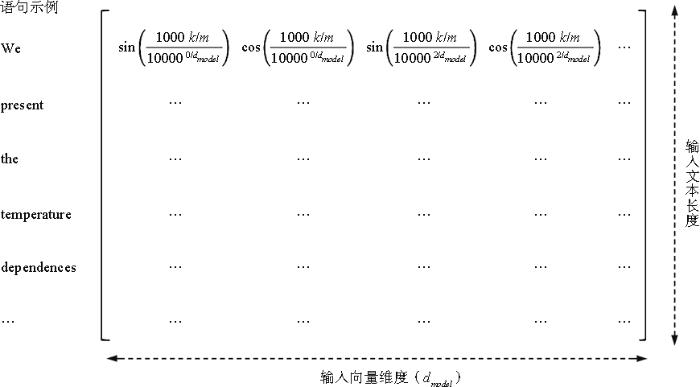
本研究基于Transformer字符位置嵌入[25]原理提出句子位置嵌入计算公式。对于篇章的第i个句子,当i为偶数时,句子位置嵌入向量元素计算方法如公式(1)所示;当i为奇数时,句子位置嵌入向量元素计算方法如公式(2)所示。
其中,
图4
3.2 多层感知机分类器
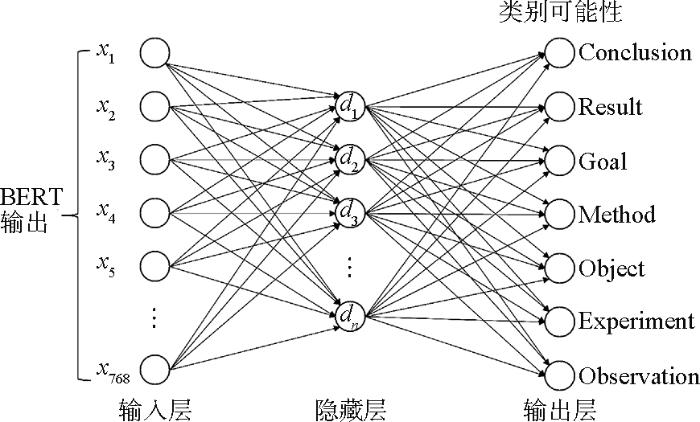
如图5所示,本文分类器的原理是在BERT最后一层隐含状态的输出后增加多层感知机,在输出层通过softmax激活函数对论文句子实现语步分类。多层感知机结构为一个全连接隐藏层及输出层。
分类器的输入,即BERT模型的<CLS>位置对应输出,为768维的向量。中间隐藏层和输出层为全连接层。隐藏层节点数n采用256、128、64三种进行实验。输出层节点数量由分类类别数确定,按照分类任务为11个和7个。输出模型的损失函数采用交叉熵损失(Cross Entropy Loss)函数。训练过程中,BERT模型和最后全连接分类层参数联合调优,实现分类模型的收敛。
图5
图5
多层感知机论文语步分类器结构
Fig. 5
Multilayer Perceptron Classifier for Argumentative Zoning
3.3 预训练模型
本文模型的初始参数采用SciBERT[26]预训练参数。该参数基于114万篇学术论文全文训练。82%的语料来自生物医学领域,与本研究的数据集领域相近,适用于该分类任务。模型深度神经网络为12层(Transfomer块),隐藏状态尺寸为768,自注意力头数为12,一共有1.1亿个参数。
SciBERT预训练模型的大部分参数将保持不变。在图2所示模型上,采用实验数据进行训练,分类器以及BERT模型的最后两层网络的参数将进行优化,实现损失函数的最小化。
4 实验过程
4.1 数据来源
表1 ART Corpus数据集论文语步分类类别
Table 1
| 类别 | 类别缩写 | 中文含义 |
|---|---|---|
| Conclusion | CON | 结论 |
| Result | RES | 结果 |
| Goal | GOA | 目标 |
| Method | MET | 方法 |
| Object | OBJ | 对象 |
| Experiment | EXP | 实验 |
| Observation | OBS | 观察 |
| Hypothesis | HYP | 假设 |
| Motivation | MOT | 动机 |
| Background | BAC | 背景 |
| Model | MOD | 模型 |
4.2 数据预处理
图6
表2 各语步标签的论文语句统计
Table 2
| 统计指标 | CON | RES | GOA | MET | OBJ | EXP | OBS | HYP | MOT | BAC | MOD |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 句子数 | 3 082 | 7 349 | 548 | 3 740 | 1 189 | 2 822 | 4 643 | 655 | 465 | 6 648 | 3 449 |
| 占比(%) | 8.91 | 21.25 | 1.58 | 10.81 | 3.44 | 8.16 | 13.42 | 1.89 | 1.35 | 19.22 | 9.97 |
| 平均单词数 | 28.10 | 26.70 | 28.46 | 25.07 | 25.16 | 24.33 | 22.81 | 27.33 | 25.39 | 25.50 | 27.16 |
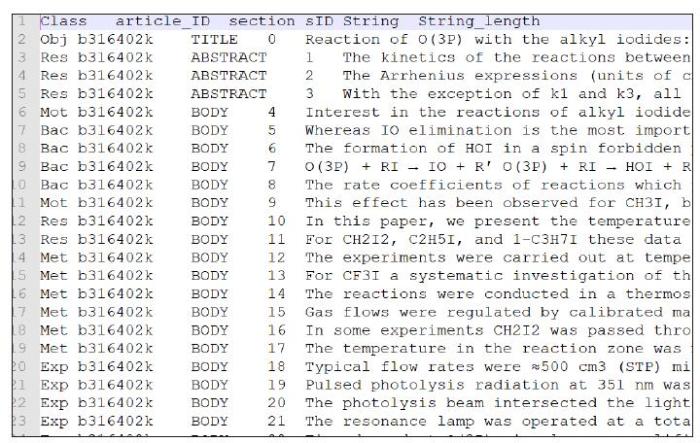
数据中已标注句子所属(标题、摘要、正文),以及句子序号。由于标题、摘要与正文句子的重要性和位置属性不同,在处理句子位置信息时,区分标题、摘要、正文。分别将标题、摘要、正文作为单独篇章,按照3.1所述方法计算各句子位置嵌入向量,用于后续处理。
为验证分类模型的健壮性,在上述数据格式转换和噪声数据清理后,进行数据筛选,过滤出前7种核心语步标签,形成只包含前7个类别的数据子集。
4.3 实验方法
实验分为两部分:第一部分采用全部11个标签的数据集;第二部分仅采用前7个核心分类的数据集,以验证在核心语步分类上的效果。两部分实验分别采用本文改进的模型输入与原始的模型输入进行分类结果比较。以篇章为单位随机选取实验训练集、开发集、测试集,比例分别为82%、8%、10%。
由于模型深度和参数数量较大,训练需要较好的硬件支持。本文模型采用Python 3.5语言,基于TensorFlow 1.13和Google BERT开源代码开发。硬件环境为Intel Xeon 16核处理器、Nvidia Tesla P100显卡、64GB内存。
5 实验结果及分析
实验结果从准确率(Accuracy)、召回率(Recall)、F1三个维度进行评价。在上述实验环境下,单个模型的训练时间约需一个小时。改进输入的模型分类任务的最佳模型超参数如表3所示。
表3 各分类任务的最佳模型超参数
Table 3
| 分类模型 | 批处理大小 | 学习率 | 训练期 | 分类器隐含层节点数 |
|---|---|---|---|---|
| 11标签分类 | 16 | 4 | 256 | |
| 7标签分类 | 32 | 4 | 128 |
基于同样的实验数据,分类模型效果对比如表4所示。本文分类总体准确率较支持向量机模型LibSVM(51.6%)提高了29.7%,达到81.3%;平均召回率及平均F1均大幅提高。
表4 语步分类结果对比
Table 4
| 分类模型 | 总体 准确率(%) | 平均 召回率(%) | 平均F1(%) | |
|---|---|---|---|---|
| LibSVM 11标签 | 51.6 | 43.0 | 46.3 | |
| 11标签分类 | SciBERT | 75.2 | 68.5 | 74.6 |
| 改进输入 | 81.3 | 72.4 | 75.5 | |
| 7标签分类 | SciBERT | 80.1 | 76.4 | 78.8 |
| 改进输入 | 85.5 | 80.7 | 83.1 | |
表4中,11标签分类以及7标签分类分别采用改进输入模型以及SciBERT模型进行对比实验。对7标签分类的核心语步数据集,本文分类模型表现出更优的语步分类能力,分类总体准确率达85.5%。改进输入的SciBERT训练模型较原模型在11标签分类和7标签分类任务上均有提高,总体准确度分别提高6.1%和5.4%。
图7
图7
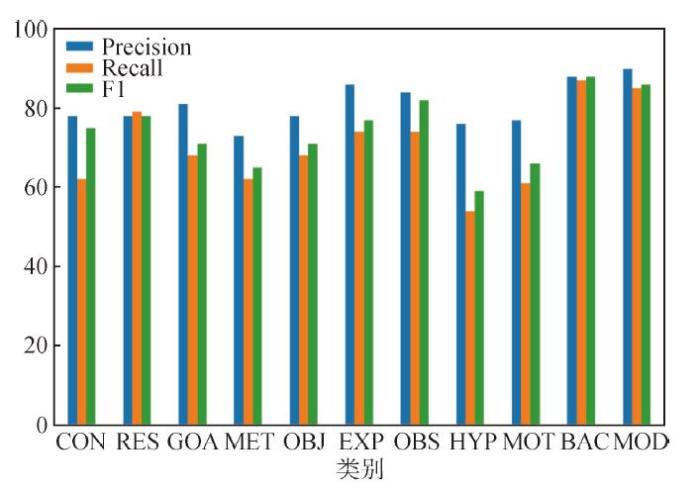
本研究模型11标签分类论文语步分类结果评价指标对比(%)
Fig. 7
Classification Metrics on 11-class Argumentative Zoning for Each Class(%)
如图7所示,11标签分类任务结果表明对“实验”(EXP)、“背景”(BAC)、“观察”(OBS)、“模型”(MOD)4类语步分类效果最好,F1值分别为78.7%、88.3%、81.6%、86.2%。从训练数据的分布分析,“背景”和“观察”语步在训练集中占比较高(分别为19.22%和13.42%),可解释较好的分类效果。但“实验”和“模型”语步在训练集中占比较低(分别为8.16%和9.97%),却获得较好的分类效果,这可能与这两个语步显著的化学实验描述专业写作特点(如大量的化学反应式等)有较强关系。
6 结论
本文将深度学习语言表征模型与学术文献语步分类任务结合,改进模型输入,增加表征句子位置的输入,对模型进行迁移学习和联合调优。该方法可学习单词潜在的上下文语境特征,生成句子级别的嵌入表达,避免了传统分类方法的特征构建步骤。模型训练基于预训练模型参数进行迁移学习,可大幅缩短模型训练时间,同时保证模型效果。此外,该模型不受限于语言类别,可适用于各种语言的学术论文语类语篇研究。
采用公开的数据集进行模型训练、测试评估。实验结果表明,该方法较传统的“特征构建+机器学习”分类模式的分类效果有较大提高。此外,由于句子位置能在一定程度上反映论文语步属性,结合句子位置信息的模型输入对模型效果有一定程度的提高。基于以海量学术论文语料训练的预训练模型SciBERT,本提出的改进模型输入的方法,模型分类效果较浅层学习方法有大幅提高,较原SciBERT模型也有一定程度提高,具有实际应用潜力。受限于实验环境,本文提出的模型输入基于预训练参数训练,模型参数尚有进一步拟合空间。
作者贡献声明
王末,崔运鹏:提出研究思路,设计研究方案,进行实验,论文起草;
陈丽,李欢:采集、清洗数据;
王末:论文最终版本修订。
利益冲突声明
所有作者声明不存在利益冲突关系。
支撑数据:
支撑数据见期刊网络版,
[1] Liakata Maria, Soldatova Larisa.ART_Corpus.tar.gz. The ART Corpus.
[2] 王末,崔运鹏,陈丽,李欢. preprocessed_sentence_core_concept_class.tsv. 预处理后数据集.
参考文献
Automatic Recognition of Conceptualization Zones in Scientific Articles and Two Life Science Applications
[J].
DOI:10.1093/bioinformatics/bts071
URL
[本文引用: 6]

Motivation: Scholarly biomedical publications report on the findings of a research investigation. Scientists use a well-established discourse structure to relate their work to the state of the art, express their own motivation and hypotheses and report on their methods, results and conclusions. In previous work, we have proposed ways to explicitly annotate the structure of scientific investigations in scholarly publications. Here we present the means to facilitate automatic access to the scientific discourse of articles by automating the recognition of 11 categories at the sentence level, which we call Core Scientific Concepts (CoreSCs). These include: Hypothesis, Motivation, Goal, Object, Background, Method, Experiment, Model, Observation, Result and Conclusion. CoreSCs provide the structure and context to all statements and relations within an article and their automatic recognition can greatly facilitate biomedical information extraction by characterizing the different types of facts, hypotheses and evidence available in a scientific publication.
Results: We have trained and compared machine learning classifiers (support vector machines and conditional random fields) on a corpus of 265 full articles in biochemistry and chemistry to automatically recognize CoreSCs. We have evaluated our automatic classifications against a manually annotated gold standard, and have achieved promising accuracies with 'Experiment', 'Background' and 'Model' being the categories with the highest F1-scores (76%, 62% and 53%, respectively). We have analysed the task of CoreSC annotation both from a sentence classification as well as sequence labelling perspective and we present a detailed feature evaluation. The most discriminative features are local sentence features such as unigrams, bigrams and grammatical dependencies while features encoding the document structure, such as section headings, also play an important role for some of the categories. We discuss the usefulness of automatically generated CoreSCs in two biomedical applications as well as work in progress.
Summarizing Scientific Articles: Experiments with Relevance and Rhetorical Status
[J].DOI:10.1162/089120102762671936 URL [本文引用: 2]
英语学术论文摘要语步结构自动识别模型的构建
[J].
Constructing a Model for the Automatic Identification of Move Structure in English Research Article Abstracts
[J].
A Weakly-Supervised Approach to Argumentative Zoning of Scientific Documents
[C]//
语步分析视角下的等离子体物理国际SCI期刊论文写作范式研究
[J].
Writing Paradigm of Plasma Physics SCI Journal Articles from the Perspective of Move Analysis Theory
[J].
An Annotation Scheme for Discourse-Level Argumentation in Research Articles
[C]//
英语学术语篇语类结构研究述评(1980-2012)
[J].
A Survey on English Academic Paper Genre Studies
[J].
Information Extraction from Scientific Articles: A Survey
[J].DOI:10.1007/s11192-018-2921-5 URL [本文引用: 1]
Analyzing the Dynamics of Research by Extracting Key Aspects of Scientific Papers
[C]//
Method Mention Extraction from Scientific Research Papers
[C]//
Using Argumentation to Extract Key Sentences from Biomedical Abstracts
[J].DOI:10.1016/j.ijmedinf.2006.05.002 URL [本文引用: 1]
Towards Extracting Domains from Research Publications
[C]//
Generative Content Models for Structural Analysis of Medical Abstracts
[C]//
Computational Analysis of Move Structures in Academic Abstracts
[C]//
Identifying Sections in Scientific Abstracts Using Conditional Random Fields
[C]//
Extracting Formulaic and Free Text Clinical Research Articles Metadata Using Conditional Random Fields
[C]//
Dr. Inventor Framework: Extracting Structured Information from Scientific Publications
[C]//
Mover: A Machine Learning Tool to Assist in the Reading and Writing of Technical Papers
[J].DOI:10.1109/TPC.2003.816789 URL [本文引用: 1]
Identifying the Information Structure of Scientific Abstracts: An Investigation of Three Different Schemes
[C]//
Rhetorical Move Detection in English Abstracts: Multi-label Sentence Classifiers and Their Annotated Corpora
[C]//
Automatic Argumentative-Zoning Using Word2vec
[OL].
Distributed Representations of Words and Phrases and Their Compositionality
[C]//
Glove: Global Vectors for Word Representation
[C]//
Bert: Pre-training of Deep Bidirectional Transformers for Language Understanding
[OL].
Attention is All You Need
[OL].
SciBERT: A Pretrained Language Model for Scientific Text
[C]//